Publications
2026
[1]
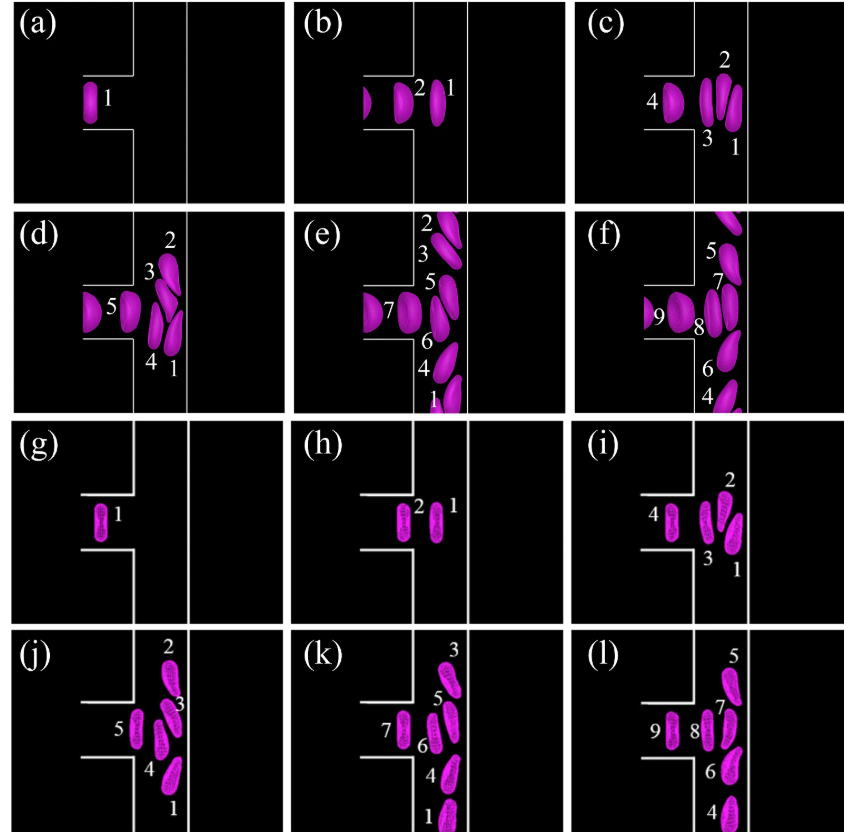
N Kadivar, G Li (equal first author)*, J Zheng, M Dao, GE Karniadakis, M Xu.
An AI-enabled tool for quantifying overlapping red blood cell sickling dynamics in microfluidic assays.
arXiv:2601.17703, 2026.
[Open Access]
2025
[2]
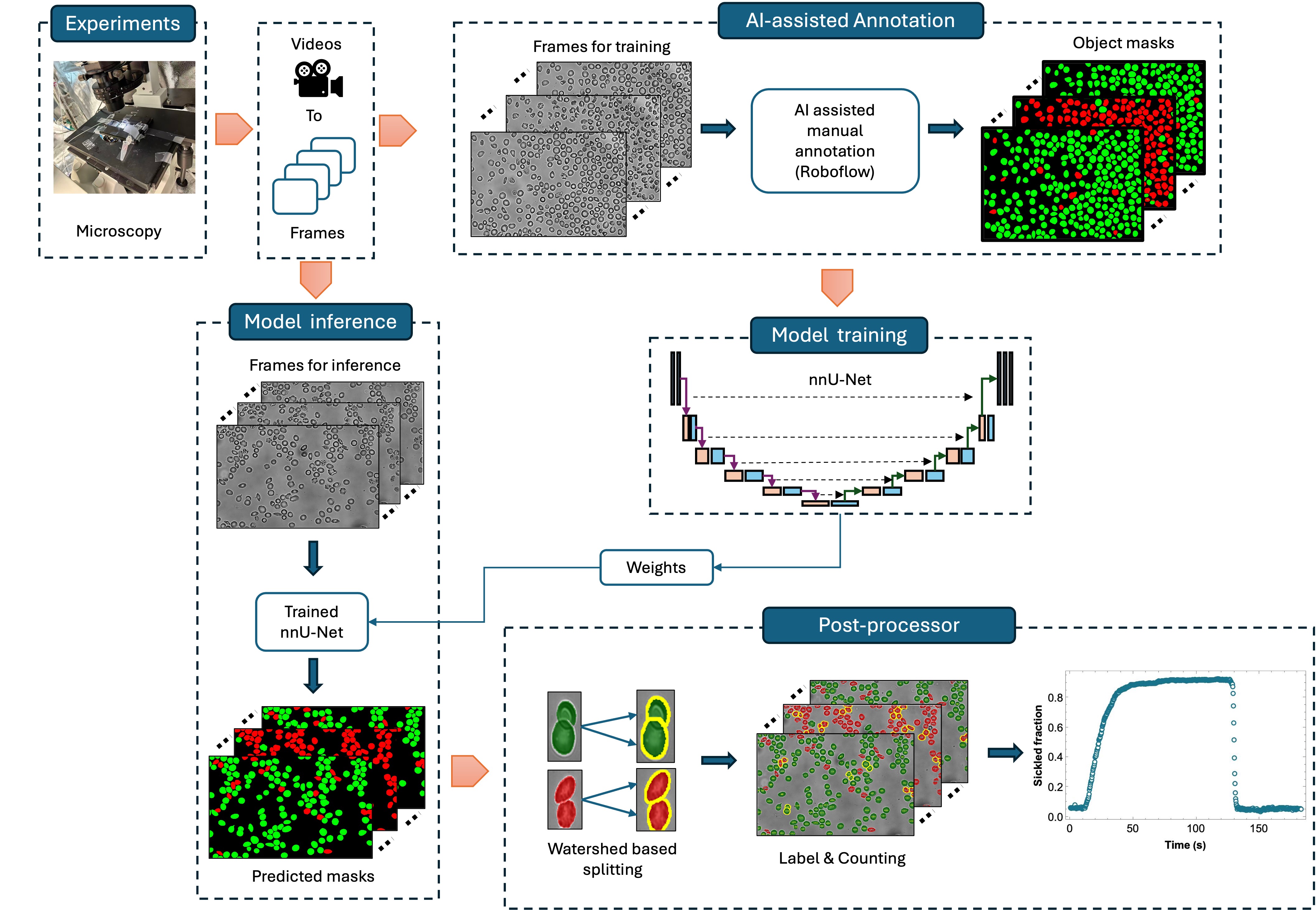
Z Chai, G Li*, PA Ndour, P Connes, PA Buffet, M Franco, GE Karniadakis.
In silico biophysics and rheology of blood and red blood cells in Gaucher Disease.
PLoS Computational Biology, 21(9): e1012705, 2025.
[Open Access]
[3]
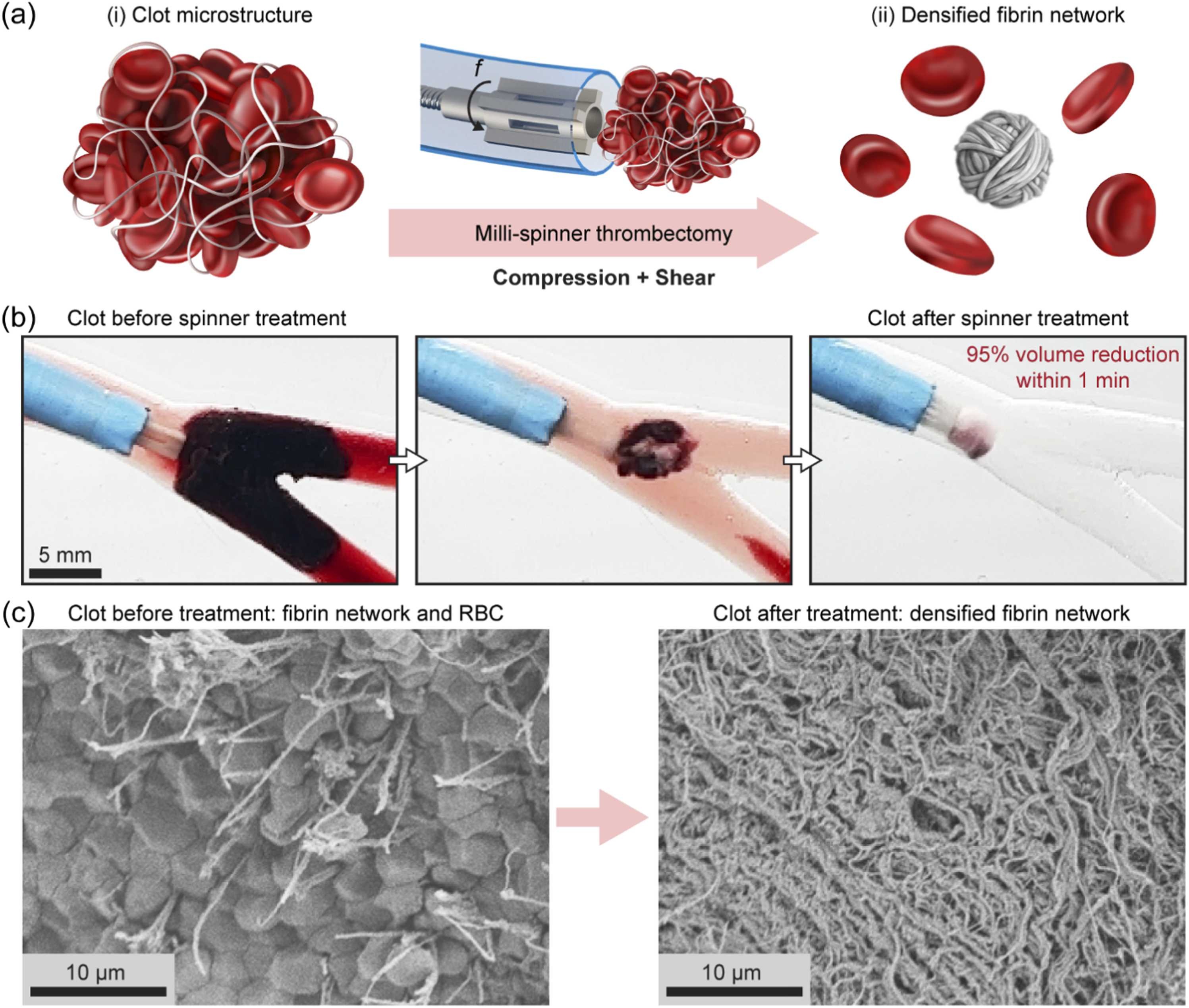
Y Chang, G Li(equal first author)*, J Sim, GE Karniadakis, RR Zhao.
Clot Treatment via Spinning-Induced Fibrin Microstructure Densification and Clot Volume Reduction.
Extreme Mechanics Letters (2025): 102391.
[Open Access]
2024
[4]
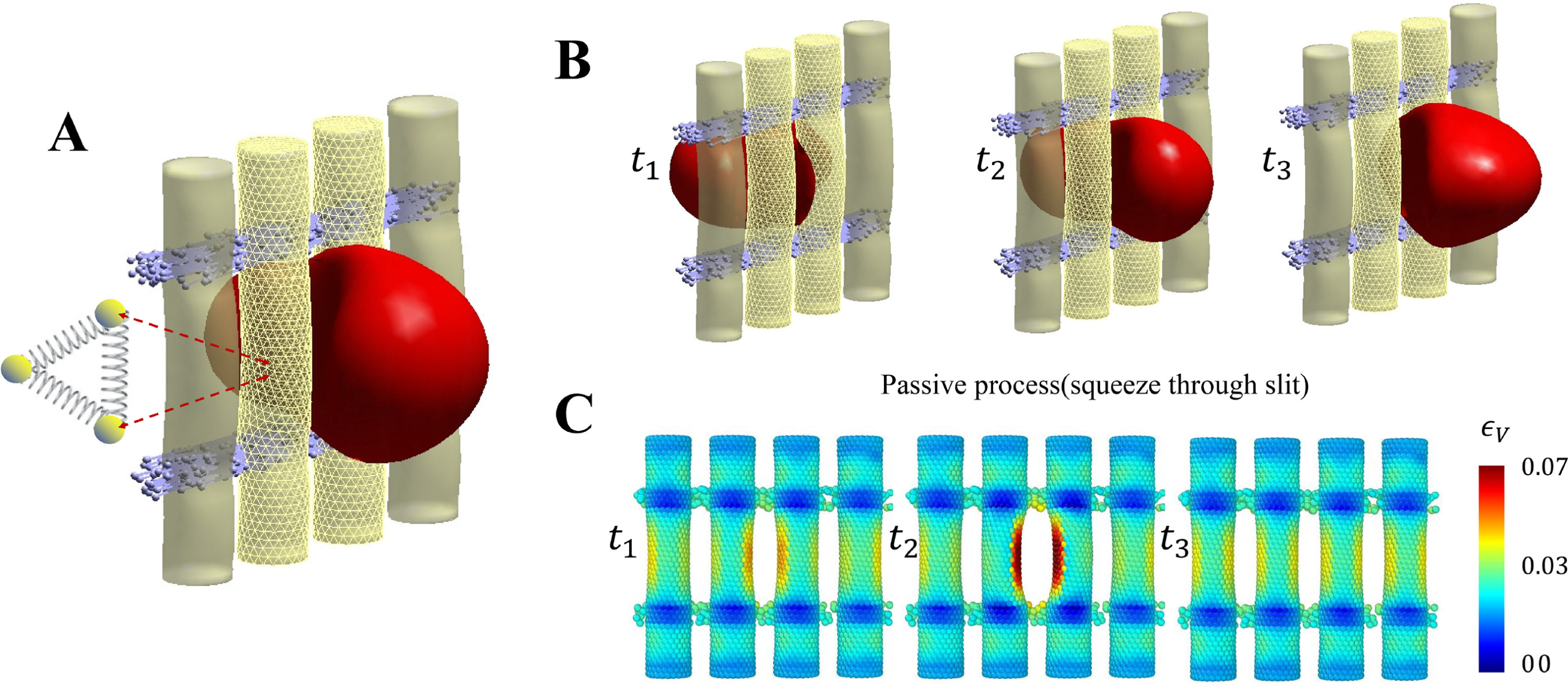
Guansheng Li*, He Li, Papa alioune Ndour, Mélanie Franco, Xuejin Li, Ian MacDonald, Ming Dao, Pierre A Buffet, George Em Karniadakis.
Red blood cell passage through deformable interendothelial slits in the spleen: Insights into splenic filtration and hemodynamics.
Computers in Biology and Medicine 182 (2024): 109198.
[Open Access]
[5]
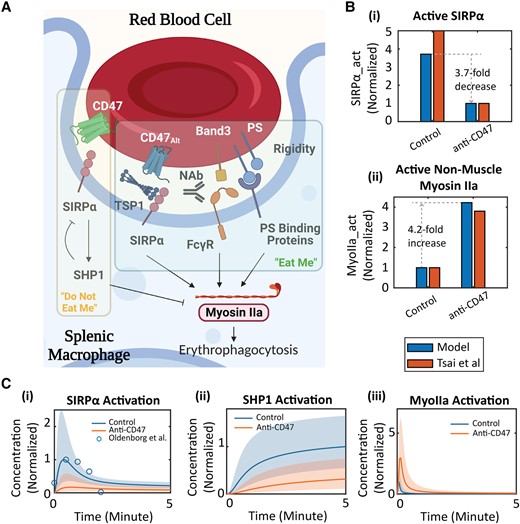
Yu Zhang, Yuhao Qiang, He Li, Guansheng Li*, Lu Lu, Ming Dao, George E Karniadakis, Aleksander S Popel, Chen Zhao.
Signaling-biophysical modeling unravels mechanistic control of red blood cell phagocytosis by macrophages in sickle cell disease.
PNAS Nexus 3.2 (2024): pgae031.
[Open Access]
2023
[6]
G Li*, Y Qiang, H Li, X Li, PA Buffet, M Dao, GE Karniadakis.
A combined computational and experimental investigation of the filtration function of splenic macrophages in sickle cell disease.
PLOS Computational Biology 19(12), e1011223.
[Open Access]
[7]
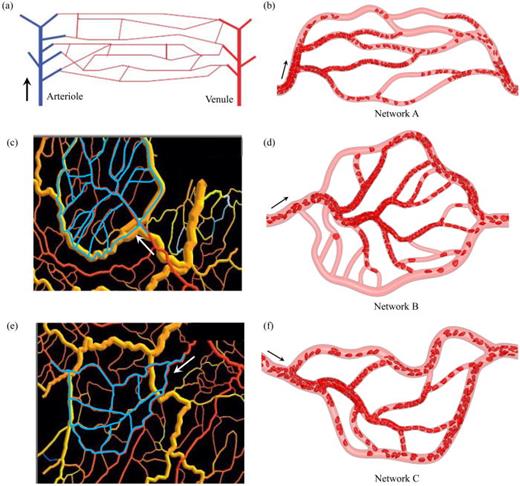
G Li*, T Ye, Z Xia, S Wang, Z Zhu.
Analysis and prediction of hematocrit in microvascular networks.
International Journal of Engineering Science 191, 103901.
[Open Access]
[8]
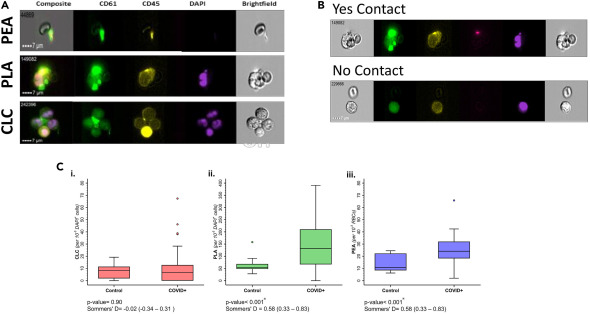
Ander Dorken-Gallastegi, Yao Lee, Guansheng Li*, He Li, Leon Naar, Xuejin Li, Ting Ye, Elizabeth Van Cott, Rachel Rosovsky, David Gregory, Ronald Tompkins, George Karniadakis, Haytham MA Kaafarani, George C Velmahos, Jarone Lee, Galit H Frydman.
Circulating cellular clusters are associated with thrombotic complications and clinical outcomes in COVID-19.
iScience 26(7).
[Open Access]
[9]
G Li*, Y Qiang, H Li, X Li, M Dao, GE Karniadakis.
In silico and in vitro study of the adhesion dynamics of erythrophagocytosis in sickle cell disease.
Biophysical Journal 122(12), 2590-2604.
[Open Access]

[10]
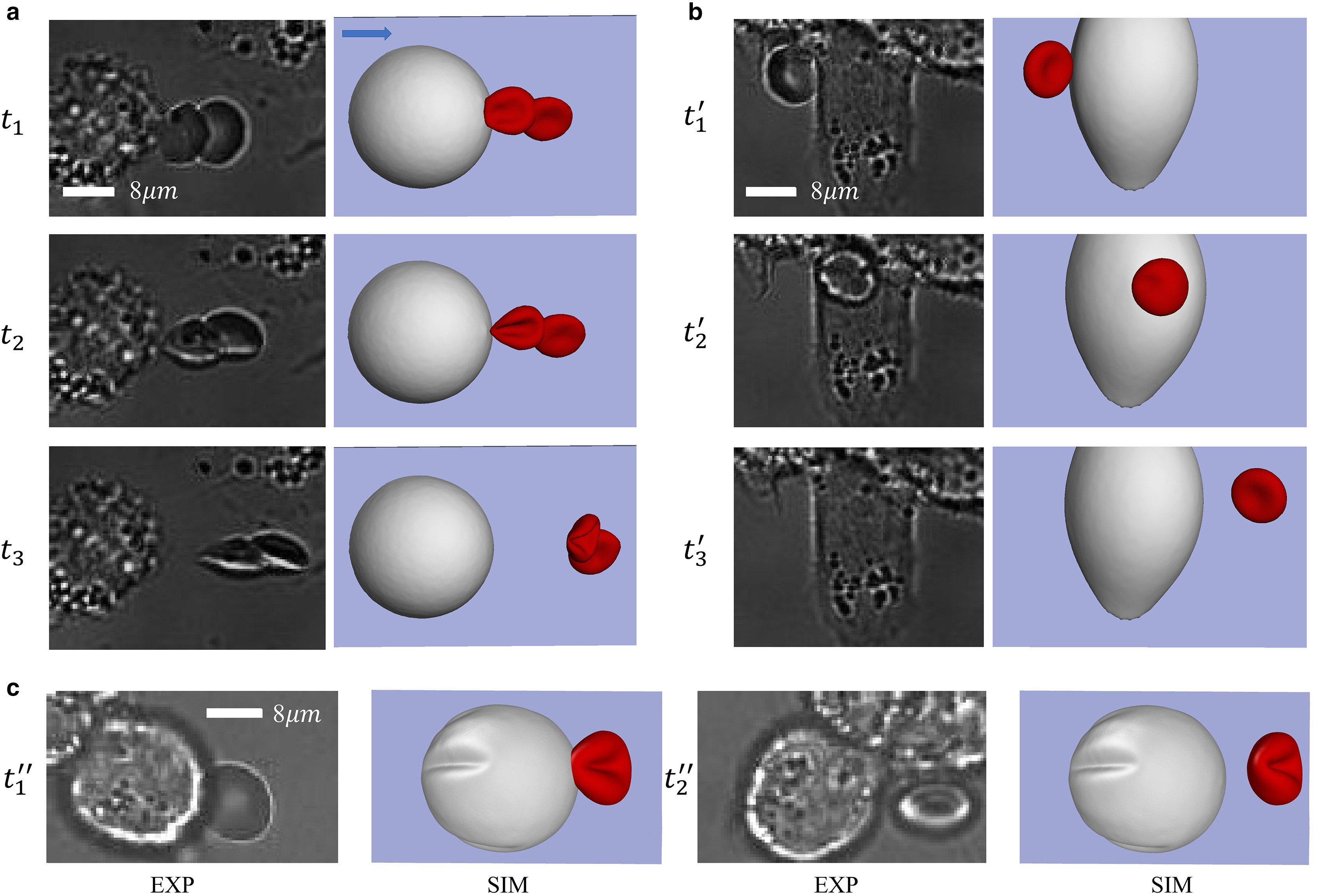
G Li*, T Ye, B Yang, S Wang, X Li.
Temporal-spatial heterogeneity of hematocrit in microvascular networks.
Physics of Fluids 35(2).
[Open Access]
2021
[11]
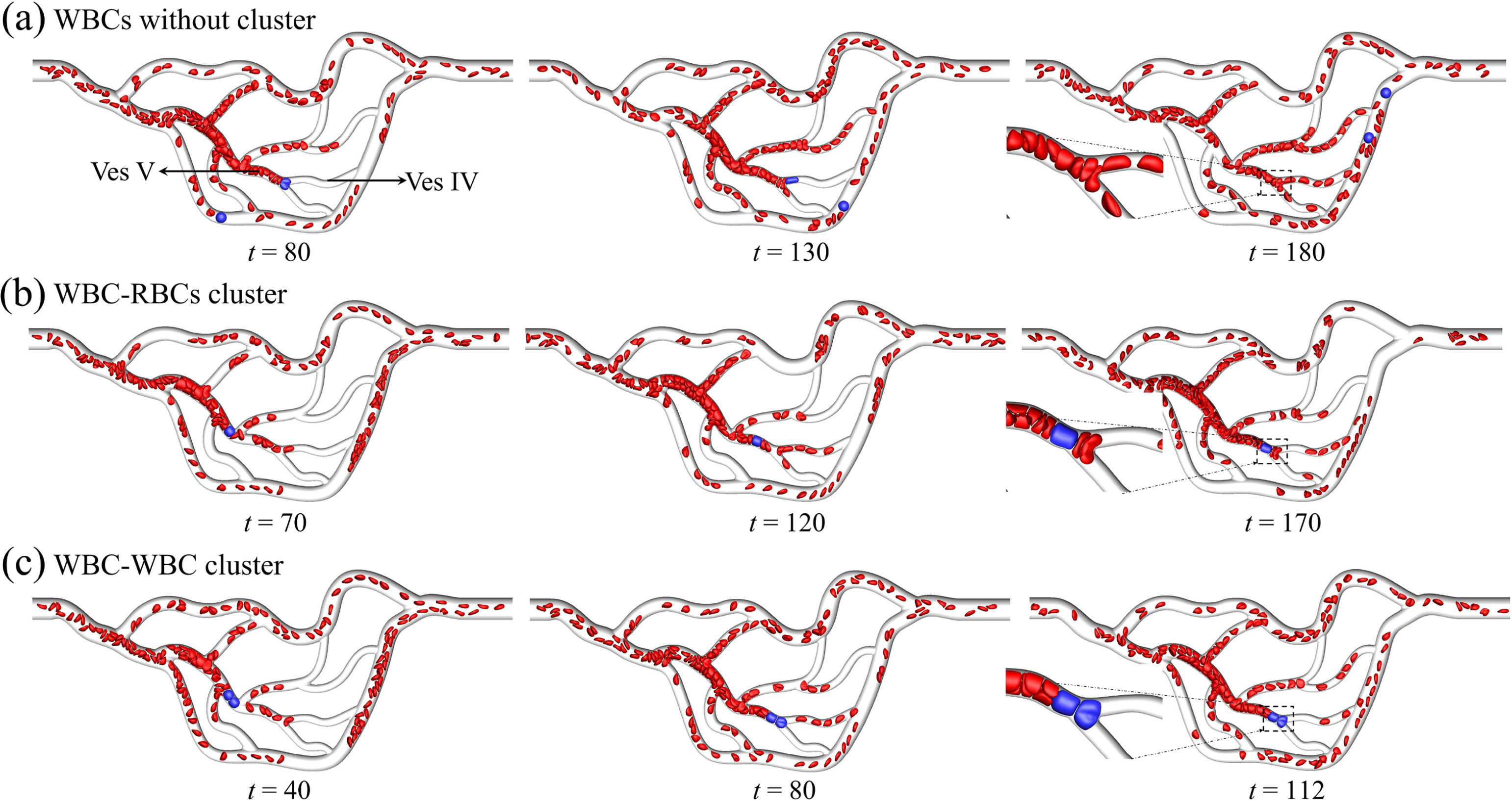
S Wang, T Ye, Guansheng Li*, X Zhang, H Shi.
Margination and adhesion dynamics of tumor cells in a real microvascular network.
PLoS Computational Biology 17(2), e1008746.
[Open Access]
2020
[12]
T Ye, X Zhang, Guansheng Li*, S Wang.
Biomechanics in thrombus formation from direct cellular simulations.
Physical Review E 102(4), 042410.
[Open Access]
[13]
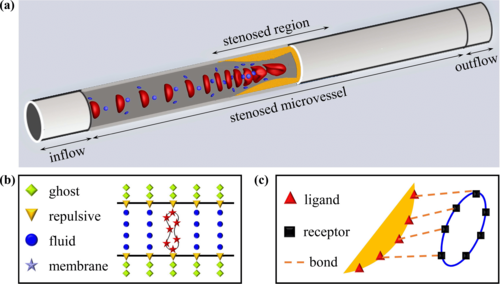
Guansheng Li*, T Ye, S Wang, X Li, R UI Haq.
Numerical design of a highly efficient microfluidic chip for blood plasma separation.
Physics of Fluids 32(3), 031903.
[Open Access]
[14]
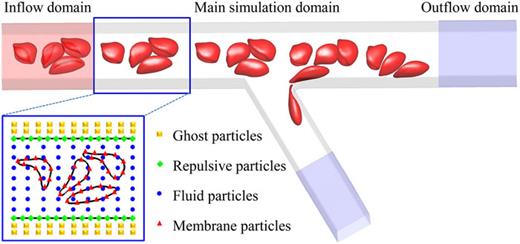
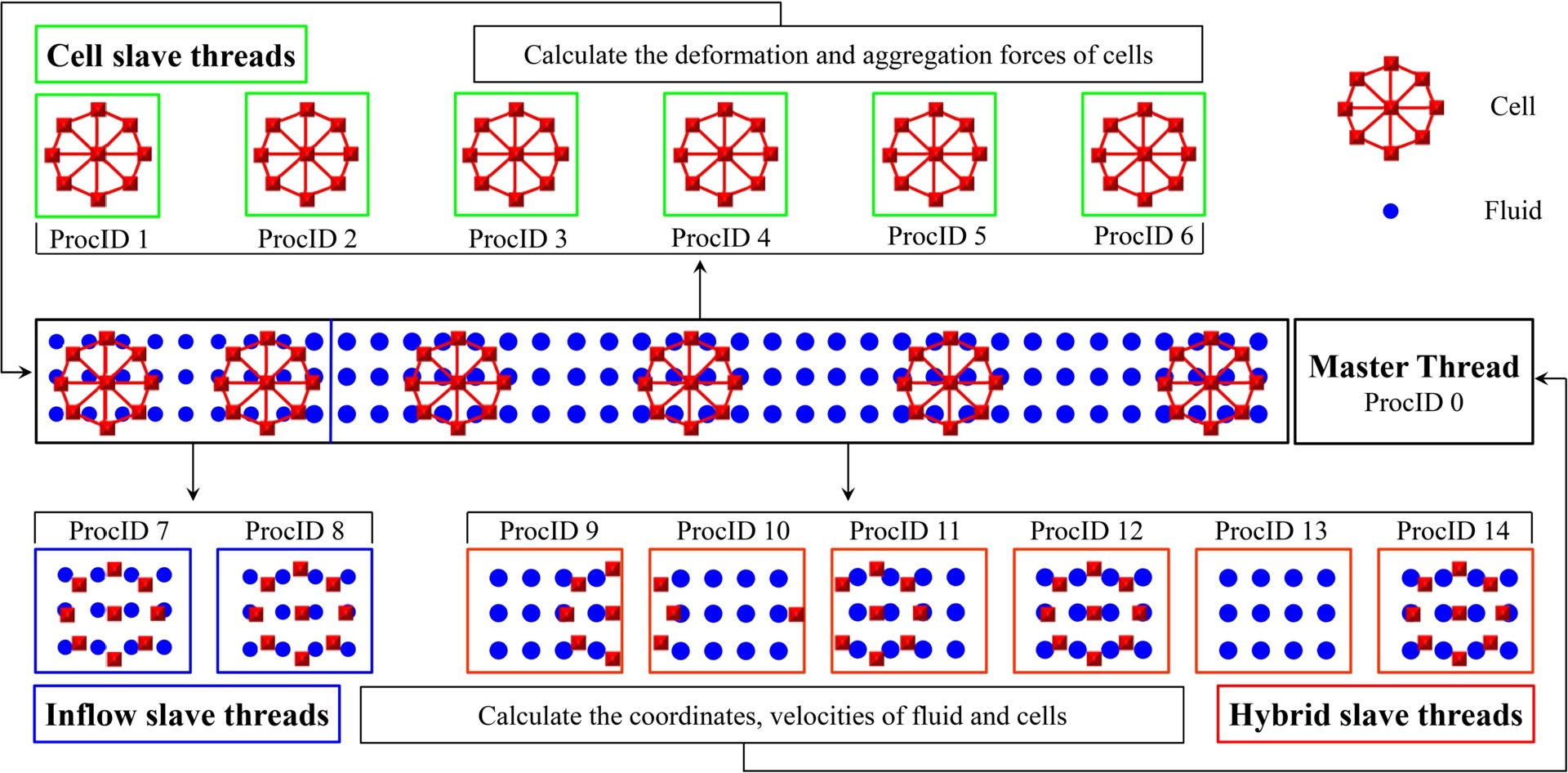
Guansheng Li*, T Ye, X Li.
Parallel modeling of cell suspension flow in complex micro-networks with inflow/outflow boundary conditions.
Journal of Computational Physics 401, 109031.
[Open Access]
2019
[15]
T Ye, L Peng, Guansheng Li*.
Red blood cell distribution in a microvascular network with successive bifurcations.
Biomechanics and Modeling in Mechanobiology 18(6), 1821-1835.
[Open Access]